Sponsored content brought to you by
The progression of next-generation sequencing (NGS) from research to clinical laboratories over the last 20 years has meant that the powerful, and long-awaited, healthcare goal of personalized medicine has started to become a reality.
There are a few technologies leveraged for NGS, with the most widely adopted being sequencing by synthesis (SBS). SBS uses labeled nucleotides that are tracked while multiple DNA chains are copied in parallel. This method is much faster and cheaper than the original Sanger sequencing method that was used in the Human Genome Project and makes large-scale whole-genome sequencing (WGS) more accessible.
From a clinical standpoint, NGS has opened up a whole host of opportunities to better understand disease pathology and risk factors. It has had a big impact on cancer where it can be used to identify mutations that might make tumors more (or less) susceptible to certain treatments. Since 2006, The Cancer Genome Atlas Program, a collaboration of 20 institutions across the United States and Canada, has mapped key genomic changes in 33 different tumor types including 10 rare cancers.1 The information generated by the project has improved understanding of the molecular basis of cancer, revolutionized how cancer is classified, and identified therapeutic targets.
Other clinical NGS applications include the study and diagnosis of rare diseases, noninvasive prenatal testing, infection biology, and pharmacogenomics, where medicines are being prescribed based on a patient’s genetics. NGS is also being used in population-based studies to create polygenic risk scores that will predict future risk for common diseases like diabetes, dementia, and heart disease.

The Impact of in-vitro diagnostic regulations
Up to now, the majority of this work has been carried out in a research setting. However, as countries begin to implement NGS sequencing into clinical practice, the focus shifts to tighter regulatory approval since the instruments and their associated reagents and assays will fall under the umbrella of in-vitro diagnostic medical devices (IVDs).
The U.S. Food and Drug Administration (FDA) defines IVDs as “reagents, instruments, and systems intended for use in diagnosis of disease or other conditions, including a determination of the state of health, in order to cure, mitigate, treat, or prevent disease or its sequelae2.” They say “[s]uch products are intended for use in the collection, preparation, and examination of specimens taken from the human body2.” The FDA regulations classify IVD products into three groups according to the degree of regulatory control needed to “reasonably assure safety and effectiveness2.”
In Europe, new IVD regulations were introduced in 2017, with the transitional period for implementation ending in May this year. The change has led to the proportion of IVDs needing certification by a notified body to increase from around 20% to approximately 80%3. This is because the previous regulatory classification was based on a set list of conditions or pathogens that a given diagnostic medical device was designed to diagnose. If a pathogen was not on the list—as was the case for SARS-CoV-2—diagnostic tests for that pathogen fell into the lowest risk category and did not need to be verified by a regulatory agency4. The new regulations are based on risk rather than a set list. Sequencers for NGS applications along with library prep reagents for downstream DNA analysis by NGS sequencing come under the highest risk category (class A)5. This means they will undergo a high level of scrutiny before reaching the market.
With the introduction of national and international genomics strategies such as WHO’s “Global genomic surveillance strategy for pathogens with pandemic and epidemic potential 2022–2032”6 and the U.K.’s “Accelerating genomic medicine in the NHS” report7, which provides a strategy for embedding genomics in the NHS over the next 5 years, an increasing number of clinical laboratories will find themselves looking for NGS sequencing solutions that can cope with higher testing rates.
High-throughput NGS on an IVD-compliant, FDA registered instrument
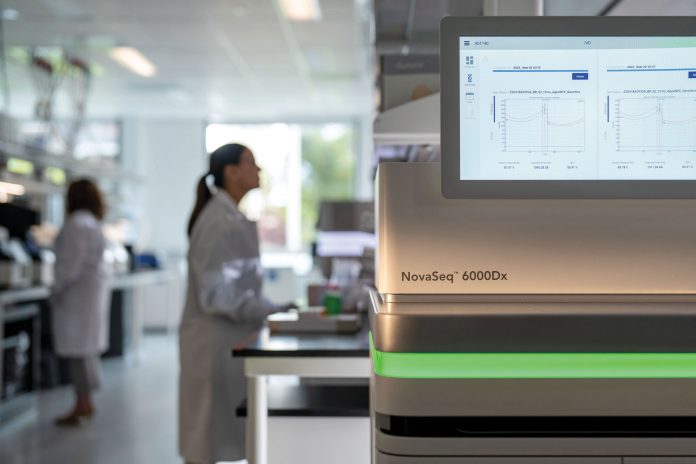
At present, Illumina is the only company to offer an FDA-registered and CE-marked high-throughput sequencing instrument. The recently launched NovaSeq™ 6000Dx comes with a new, streamlined user interface for operational simplicity and offers users the flexibility of sequencing in either IVD or research-use only (RUO) mode without the need for a reboot when switching between the two.
In IVD mode, the instrument has two different flow cell formats—S2 and S4—with maximum outputs of 1 Tb and 3 Tb, respectively, and high data quality; at least 85% of bases have Q-scores of 30 or more at read lengths of 2 x 150 bp. High quality sequencing data, specifically for variant calling, is incredibly important for therapeutically actionable insights. In addition to highly accurate data, the NovaSeq 6000Dx offers flexibility in scale and utilization. Each flow cell can be operated independently, enabling separate starts and stops and the run times are 40 hours or less with the S2 format and 44 hours or less with S4.
One of the key diagnostic applications of the NovaSeq 6000Dx is target enrichment. The regulatory-approved Illumina DNA Prep with Enrichment Dx kit comprises a set of reagents and consumables that are used with the laboratory’s own probe panels to prepare sample libraries from genomic DNA derived from human cells and tissue.
Another key feature is assay development, and in the future, Illumina expects to offer a wider portfolio of their own IVD assays for the NovaSeq 6000Dx as well as those developed with commercial partners. When in RUO mode, the instrument can also be used for whole-genome sequencing, whole-transcriptome sequencing, targeted RNA Sequencing such as exome and custom enrichment panels, and methylation sequencing.
Streamlined data handling
As laboratories begin to increase the volumes of sequencing they are carrying out, the amount of complicated genomic data they produce also increases along with the need for dedicated bioinformaticians to interpret such data.
To simplify this process, Illumina has paired a DRAGEN™, short for Dynamic Read Analysis for GENomics, server with each NovaSeq 6000Dx. The DRAGEN platform streamlines data analysis by carrying out BCL Conversion, FASTQ generation, mapping/aligning, and variant calling all within one module. If the DRAGEN ORA compression option is enabled, it will also automatically compress FASTQ files, which results in an approximate fivefold reduction in file sizes, faster file transfers, and reduced data storage costs.
Taken together, this means that the NovaSeq 6000Dx instrument is part of an integrated, three-step workflow that includes library preparation, high-throughput sequencing in either IVD or RUO Mode, and accelerated secondary data analysis with a paired DRAGEN server. The increased efficiency associated with the higher number of samples per run relative to lower throughput instruments translates to large reductions in the price per genome—another important consideration when clinical laboratories begin to scale-up their NGS offerings.
As an increasing number of clinical laboratories find themselves looking for NGS sequencing solutions that tick all the boxes, the NovaSeq 6000Dx should surely be high up on the wish list.
References
- www.cancer.gov/about-nci/organization/ccg/research/structural-genomics/tcga/history [accessed November 1, 2022]
- www.fda.gov/medical-devices/ivd-regulatory-assistance/overview-ivd-regulation [accessed November 1, 2022]
- www.tecan.com/blog/how-to-navigate-the-ivdd-ivdr-transition-part-1 [accessed November 1, 2022]
- qbdgroup.com/en/blog/ivdr-classification-of-in-vitro-diagnostic-medical-devices-guide [accessed November 1, 2022]
- health.ec.europa.eu/system/files/2022-01/md_mdcg_2020_guidance_classification_ivd-md_en.pdf [accessed November 1, 2022]
- Carter LL, Yu MA, Sacks MA et al. Bull World Health Organ 2022; 100 (4): 239–239A.
- www.england.nhs.uk/long-read/accelerating-genomic-medicine-in-the-nhs [accessed November 1, 2022]
For In Vitro Diagnostic Use. Not available in all countries or regions.