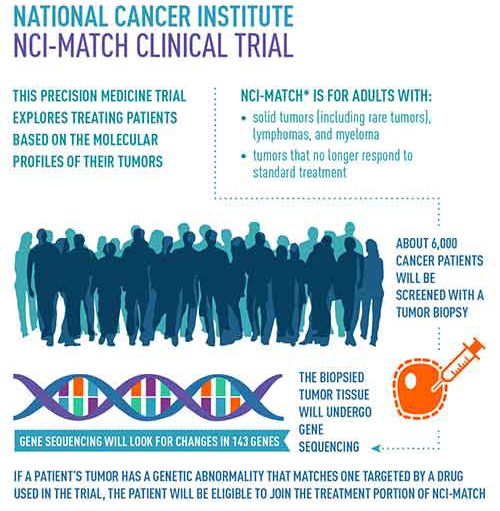
In the fight against cancer, scientists and clinicians have looked toward novel approaches that provide a more comprehensive understanding of the molecular mechanisms that drive the disease—all with the hope of uncovering new therapies that afford many patients a better quality of care. To that end, in August of 2015, the National Cancer Institute (NCI) began enrolling patients into a unique initiative that looked to analyze cancer patients’ tumors for actionable mutations for which a targeted therapy already exists. The unique aspect is that patients were grouped based on their mutations and not their specific cancer—with the hope that treating these cancers on the basis of their mutational profile will be more effective. The program set up by NCI is called the Molecular Analysis for Therapy Choice (NCI-MATCH) and is looking to examine tumor samples from approximately 6,000 patients.
The goals are ambitious, and complicated by the concerns expressed by many researchers about the complexity, accuracy, and reproducibility of next-generation sequencing (NGS) for application in clinical trials. However, investigators involved with NCI-MATCH set out to address whether the methodologies were valid that the clinical centers were using to isolate, sequence, and analyze the array of tumor samples being collected.
Now, a recent report in The Journal of Molecular Diagnostics—entitled “Analytical Validation of the Next-Generation Sequencing Assay for a Nationwide Signal-Finding Clinical Trial”—confirms that the assay tailored for this trial is highly sensitive for detecting genetic mutations from a variety of tumor tissue and, for the first time, has been reproduced with accuracy by multiple clinical laboratories, laying the groundwork for future clinical utility. The results clearly show that locked and controlled procedures permit the reliable, accurate, and reproducible use of NGS for clinical purposes.
The NCI-MATCH investigators, using the Oncomine Cancer Panel assay and the Personal Genome Machine, were able to detect more than 4,000 predefined genomic variations across 143 genes, including single nucleotide variants (SNVs), insertions/deletions (indels), copy number variations (CNVs), and gene fusions.
“We report the analytical validation processes for the NGS assay that was tailored for regulatory compliant use in the trial,” the authors wrote. “The Oncomine Cancer Panel assay and the Personal Genome Machine were used in four networked laboratories accredited for the Clinical Laboratory Improvement Amendments (CLIA). Using formalin-fixed, paraffin-embedded clinical specimens and cell lines, we found that the assay achieved an overall sensitivity of 96.98% for 265 known mutations and 99.99% specificity. High reproducibility in detecting all reportable variants was observed, with a 99.99% mean interoperator pairwise concordance across the four laboratories.”
186 samples and 12 cell lines were tested at four different laboratories. Steps were taken to maximize standardization, including the development of standard operating procedures, use of the same commercial assay and instruments, and face-to-face discussions. The investigators found that the assay results were highly reproducible, with the same results across multiple laboratories. This is critical for future clinical use leading to improved patient outcomes.
“This analytical validation study clearly found that the assay met the expected performance requirement for the intended use,” the authors penned. “This validation effort indicates that NGS assays can be robust and reproducible if the assay system is defined and standardized by locked SOPs. The process described in this article can serve as a template for other investigators who develop and validate NGS assay systems.”
Most importantly, the authors noted that NCI-MATCH NGS assay was able to accurately determine genetic abnormalities in biopsies from the pancreas, melanoma, bone, and skin samples, which “suggests that nucleic acid specimens recovered from multiple tumor tissue types are acceptable.”
“The validation study reported by Williams and colleagues is another step moving the field closer to the time when precision medicine will generate the expected benefits in improved clinical outcomes,” concluded Elizabeth Unger, M.D., Ph.D., chief of the Chronic Viral Disease Branch (CVDB), of the Division of High-Consequence Pathogens and Pathology at the Centers for Disease Control and Prevention. “Although the success of the NCI-MATCH trial cannot be assured, linking precision laboratories to precision medicine trials assures that data used for drug assignment will be reliable. Further, the use of a commercial platform and integrated analysis and reporting pipeline will greatly facilitate the broader translation of any successes.”
To see more articles from the March/April issue of Cliniclal OMICs click here.