While amplified RNA tests are the standard tests for COVID-19, the need for RNA extraction and amplification poses some challenges, namely, expensive reaction steps and equipment and limited scaling.
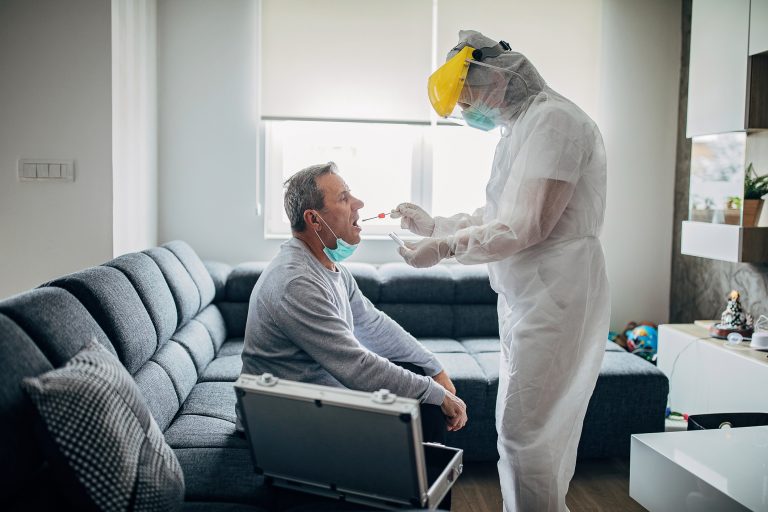
Researchers at the Karolinska Institute in Sweden have developed a test that bypasses those limitations, apparently without sacrificing accuracy. The test allows quicker, less expensive tests to be conducted in clinical as well as non-clinical settings thereby extending access to COVID-19 testing.
Currently, commercial RNA-based COVID-19 tests require samples to undergo RNA extraction to access the virus’s genetic material for diagnosis. The new test skips traditional RNA purification steps. Instead, the patient sample is heated to inactivate any virus, and moves directly to the diagnostic step. Another novelty involves the mix of chemicals in the collection and transport medium. Once the patient sample is placed in the transport medium and the sample is heated, chemicals in the buffer break it down, making the viral RNA available for immediate testing. The protocol is published in a paper in the journal Nature Communications.
“By replacing the collection buffer with simple and inexpensive buffer formulations, we can enable viral detection with high sensitivity directly from the original clinical sample, without any intermediate steps,” says senior author Björn Reinius.
The researchers compared their SARS-CoV-2 RNA-extraction-free single-reaction RT-PCR testing on 597 heat-inactivated nasopharyngeal swab samples in transport medium and compared the results with clinically diagnosed patient samples. Those samples had previously been tested following RNA extraction and traditional RT-PCR methods.
First, the team determined that the recommended heat activation protocol was 95–98 °C between 5-15 minutes. Next, they tested whether hid-RT-PCR could be improved by addition of the thermostable chemical, RNase inhibitor polyvinylsulfonic acid (PVSA), and/or the chelating agent ethylenediaminetetraacetic acid (EDTA) during heat inactivation.
The team ranked the treatments within each sample and day and observed that 95 °C 5 min + 150 μg/ml PVSA was optimal, followed by 95 °C 5 min without additives across day 0 to 4. The benefit of EDTA and higher concentrations of PVSA only became apparent after 4–7 days in storage (4 °C).
Since heat inactivation cleaves RNA into shorter fragments, the researchers tested various combinations or PCR prime and probe lengths to yield more intact RNA strands. They observed that the primer-probe set with the shortest amplicon (72 bp) performed best in hid-RT-PCR and the longest amplicon (113 bp) performed the worst.
Once the optimal heat-inactivation/buffer solution was identified, they tested their new protocol with traditional RT-PCR. The discovered that heat-inactivated-RT-PCR (hid-RT-PCR) tested samples performed similarly as standard PCR tests. hid-RT-PCR had an accuracy, sensitivity, and specificity of 98.8% (95% confidence interval, CI95: 97.5–99.5%), 96.0% and 99.8%, respectively.
From limited experiments, the researchers hypothesized that the direct RT-PCR pipeline for COVID-19 testing might also be implemented by sampling directly into a lysis buffer containing detergents such as Triton X-100. Lysed samples could immediately be subjected to RT-PCR analysis and diluted in the RT-PCR master mix without intermediate steps.