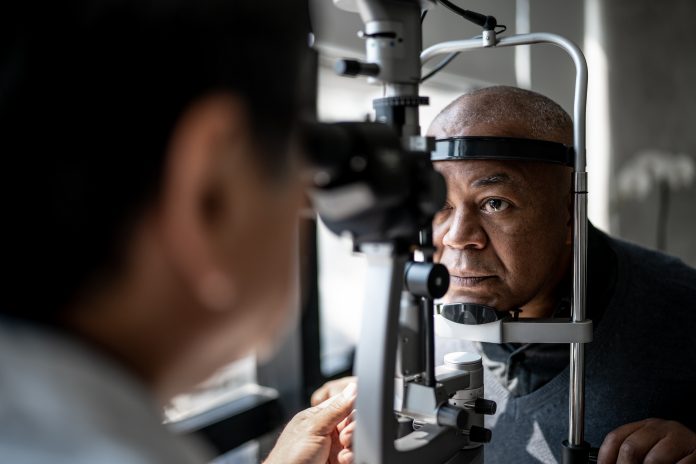
Researchers at the UCL Institute of Ophthalmology and Moorfields Eye Hospital have developed an artificial intelligence (AI) technology that can predict which genes have pathogenic variants based on a retinal scan in individuals with inherited retinal diseases.
Notably, the technology—Eye2Gene—can make these predictions without the need for human phenotype ontology (HPO) terms, which are commonly used by programs such as the Exomiser to select the most relevant disease-related genes based on known genotype phenotype relationships. Identifying such terms can be difficult and time intensive and requires specific expertise.
Nikolas Pontikos, a group leader at the UCL Institute of Ophthalmology and Moorfields Eye Hospital, presented Eye2Gene and his group’s initial findings with the tool at the European Society of Human Genetics annual conference in Glasgow this week.
He explained that inherited retinal diseases impact around 30,000 people in the U.K. and are diverse and caused by more than 270 different gene mutations. “It’s the leading cause of blindness in the working age population, over 40% of the patients remain undiagnosed, and it’s a significant health economic burden because this is a lifelong condition,” he emphasized.
The routine time to a genetic diagnosis is more than five years, which is problematic for a number of reasons and costs health providers and systems huge amounts of money. “The benefits of a diagnosis are significant for patients, for psychological support, prognosis, and family planning,” said Pontikos. “Also, if you have a genetic diagnosis…there are more and more gene targeted treatments.”
Genomic sequencing is now providing a new route to diagnoses for some rare diseases, including inherited retinal diseases, but overall this method still only provides answers for less than half of those affected.
After creating Eye2Gene, Pontikos and colleagues initially trained the system using data from more than 1000 patients with inherited retinal diseases. They then tested the system using information from 130 individuals with a retinal disease and a known genetic diagnosis, genome or exome data, retinal scans and in-depth HPO descriptions. The HPO gene score given to each case by the Exomiser program was compared with the Eye2Gene scores.
In over 75% of cases, Eye2Gene selected the correct gene with a score equal to or better than the HPO-based Exomiser program.
“We need further evaluation of Eye2Gene in order to assess its performance for different types of inherited retinal disease patients from different ethnicities, different types of imaging devices, and in different types of settings, for example primary vs. secondary care,” said Pontikos, in a press statement.
“Clinical trials will be required before our system can be deployed in clinics as medical device software,” he added.